- Select a language for the TTS:
- UK English Female
- UK English Male
- US English Female
- US English Male
- Australian Female
- Australian Male
- Language selected: (auto detect) - EN
Play all audios:
ABSTRACT The shells of pearl oysters, _Pinctada fucata,_ are composed of calcite and aragonite and possess remarkable mechanical properties. These shells are formed under the regulation of
macromolecules, especially shell matrix proteins (SMPs). Identification of diverse SMPs will lay a foundation for understanding biomineralization process. Here, we identified 72 unique SMPs
using liquid chromatography-tandem mass spectrometry (LC-MS/MS) analysis of proteins extracted from the shells of _P. fucata_ combined with a draft genome. Of 72 SMPs, 17 SMPs are related to
both the prismatic and nacreous layers. Moreover, according to the diverse domains found in the SMPs, we hypothesize that in addition to controlling CaCO3 crystallization and crystal
organization, these proteins may potentially regulate the extracellular microenvironment and communicate between cells and the extracellular matrix (ECM). Immunohistological localization
techniques identify the SMPs in the mantle, shells and synthetic calcite. Together, these proteomic data increase the repertoires of the shell matrix proteins in _P. fucata_ and suggest that
shell formation in _P. fucata_ may involve tight regulation of cellular activities and the extracellular microenvironment. SIMILAR CONTENT BEING VIEWED BY OTHERS PHYLOGENETIC COMPARISONS
REVEAL MOSAIC HISTORIES OF LARVAL AND ADULT SHELL MATRIX PROTEIN DEPLOYMENT IN PTERIOMORPH BIVALVES Article Open access 17 December 2020 IDENTIFICATION OF METHIONINE -RICH INSOLUBLE PROTEINS
IN THE SHELL OF THE PEARL OYSTER, _PINCTADA FUCATA_ Article Open access 27 October 2020 FUNCTIONAL SHELL MATRIX PROTEINS TENTATIVELY IDENTIFIED BY ASYMMETRIC SNAIL SHELL MORPHOLOGY Article
Open access 17 June 2020 INTRODUCTION Pearl oysters, _Pinctada fucata_, are one of the most important economical pearl production species in China and Japan and are also one of the best
studied biomineralization models1. Their shells are composed of calcite as the outer prismatic layer and aragonite as the inner nacreous layer. The biomineralized products possess superior
mechanical2 and biological properties3 compared to common calcium carbonate. The shells typically consist of 95% CaCO3 and approximately 5% organic macromolecules including proteins,
polysaccharides and lipids1. Specifically, shell matrix proteins (SMPs) play important roles in crystal nucleation, polymorphism, morphology and organization of calcium carbonate
crystallites during shell formation4. Since the cloning of the first SMP, Nacrein from _P. fucata_ in 19965, MSI606, N167, Prismalin-148, Shematrin9, lysine(K)-rich matrix protein (KRMP)9,
Aspein10, Tyrosinase11, N4012, Pif17713, Prisilkin-3914, PfN2315 and PfN4416 have been cloned and characterized. SMPs have been found to possess several functionalities: 1) they facilitate
calcite (Aspein17) or aragonite crystallization (N4012), 2) they act as framework proteins (Shematrin18 and Prisilkin-3914) and 3) they guide calcium carbonate assembly (N1619). A
comprehensive characterization of SMPs will offer an opportunity to better understand biomineralization processes and refine the current “chitin-silk fibroin gel proteins-acidic
macromolecules” model20. To achieve this characterization, proteomics has proven to be useful in identifying SMPs in high-throughput ways and has previously been used in _Lottia gigantea_21,
_Pinctada margaritifera_22, _Pinctada maxima_22, _Mytilus coruscus_23, _Acropora millepora_24, _Stylophora pistillata_25 and _Cepaea nemoralis_26. In the present study, we identified 72
SMPs from the shells of _P. fucata._ Ethylenediaminetetraacetic acid (EDTA)-extracted proteins were subjected to SDS-PAGE followed by liquid chromatography–mass spectrometry (LC-MS/MS)
analysis. Raw data from the LC-MS/MS were directly interrogated against the proteome derived from the draft genome of _P. fucata_27. Proteins with mascot scores above 5.0 and at least two
matched peptide fragments were considered to be valid and were analyzed by BLAST, SMART and InterProScan. In addition to controlling CaCO3 crystallization process proteins, proteomic
analysis suggests that diverse SMPs of _P. fucata_ contain extracellular matrix-(ECM) related proteins. Moreover, diverse domains were found, including carbonic anhydrase, Glyco_hydro_18,
Cu2_monooxygen, chitin-binding, complement control protein, von Willebrand factor type A, epidermal growth factor-like, tissue inhibitor of metalloproteinase and Laminin_G_2/3.
Immunohistological experiments showed localization of SMPs in the mantle cells, shells and synthetic calcite. Real-time PCR validated some representative genes _in vivo_. Together, our
results increase shell matrix proteins’ repertoires in _P. fucata_ and may guide the further study of SMPs. METHODS All methods were carried out in accordance with the approved guidelines.
All experimental protocols were approved by the Animal Experimental Ethics Committee of Tsinghua University, Beijing, China. SAMPLE PREPARATION The adult pearl oyster, _Pinctada fucata_
(with shells 5.5–6.5 cm in length and 30–40 g of wet weight and approximately 2 years of age) was obtained from the Zhanjiang Pearl Farm (Guangxi Province, China). In the laboratory, the
oysters were maintained at approximately 20 °C in an aquarium that contained aerated artificial seawater at 3% salinity. SHELL PREPARATION AND PROTEINS EXTRACTION Cleaned shells of _P.
fucata_ were immersed in 5% sodium hydroxide for 24 h and were subsequently rinsed in the distilled water to avoid possible contamination of soft tissues adhered to the inner surface of
nacre. The two layers of shells, the outer prismatic layer and the inner nacreous layer, were separated mechanically by abrasion before air-drying. Their fragments were pulverized (30 g) and
were then decalcified with 0.8 M ethylenediaminetetraacetic acid (EDTA, pH 8.0) for 60 h at 4 °C with continuous agitation. For extraction of the soluble matrix, the supernatant was
collected by centrifugation at 13,000 rpm for 30min at 4 °C and was then desalted by ultrafiltration (3 K). For extraction of the insoluble matrix, the above precipitation were thoroughly
rinsed with water and were treated with denaturing solution (30 mM Tris-HCl, pH 8.0, 1% sodium dodecyl sulfate (SDS), 10 mM dithiothreitol) at 100 °C for 30 min. After a short
centrifugation, the denatured samples were ready to be applied on 12% SDS-polyacrylamide gels. Proteins were stained with Coomassie Brilliant Blue and were quantified by a BCA assay kit
(Pierce). CHARACTERIZATION The morphologies of the cleaned shells were examined by scanning electron microscope (SEM) (FEI Quanta 200, 15 kV) after being sputter-coated with a thin layer of
gold nanoparticles. IMMUNOLOCALIZATION OF SMPS PRIMARY ANTIBODY PRODUCTION Rabbit polyclonal antibodies were produced by injecting mixed shell matrix proteins in New Zealand rabbits16.
WESTERN BLOTTING Proteins were electrophoretically transferred to PVDF membranes (Millipore) using a Mini Trans-Blot® (Bio-Rad). Then, the PVDF membranes were blocked with 5% skim milk and
were incubated with primary antibody (1:4000) for 2 h. After washing with Tris-Buffered Saline with 0.05% Tween 20 (TBST) and incubating with HRP-conjugated goat anti-rabbit secondary
antibody (1:10000, Huaxingbio Science, China), detection was performed using 3,3’-diaminobenzidine (DAB) solution (TIANGEN, China). A control experiment was performed without the first
antibody step. IMMUNOLOCALIZATION ON THE SHELLS Immunogold-labeling assays were conducted as described by the literature15,28 with some modifications. The antibodies were used at dilutions
of 1:200. Goat anti-rabbit antibodies coupled to 15 nm gold particles (1:100, Huaxingbio Science, China) were used as the secondary antibodies. A control experiment was performed without the
first antibody step. For high-resolution imaging, samples were sputter-coated with carbon and were analyzed using a Hitachi-SU8010 SEM at backscatter mode. IMMUNOLOCALIZATION ON THE MANTLE
CELLS Deparaffinized 10 μm sections of the mantle tissues, previously fixed for 24 h in Davidson fixative, were permeabilized for 10 min in TBST. Tissues were then incubated for 1 h in
saturating medium (1% BSA, TBS) at room temperature. Then, samples were incubated with the anti-matrix antibody (1:100) for 1 h in TBST–BSA 1% at room temperature (RT). After rinsing in
saturating medium, samples were incubated for 2 h at RT with HRP-conjugated goat anti-rabbit secondary antibody (1:10000, Huaxingbio Science, China). Finally, samples were observed with a
DM-4000B Leica microscope. A control experiment was performed without the first antibody step22. PROTEOMIC ANALYSIS Protein bands selected from ESMs and EISMs of the prismatic and nacreous
layers were excised and completely destained by washing with 50 μL of 50 mM NH4HCO3/CH3CN (50/50) mixture for 30 min at 37 °C. Then, reduction was conducted with 50 μL of 10 mM DTT in 50 mM
NH4HCO3 for 1 h at 57 °C and alkylation was performed with 50 μL of 100 mM iodoacetamide (IAA) for 30 min at RT in the dark. The cut gels were dried in CH3CN and were treated with 0.4 μg
trypsin (Proteomics grade, Sigma) in 10 μL of 50 mM NH4HCO3 for 12 h at 37 °C. The solution was treated with 50 μL of 1% formic acid at 30 °C for 30 min under agitation. The digests were
then lyophilized and suspended in 30 μL of 0.1% trifluoroacetic acid (TFA) and 4% acetonitrile for LC–MS/MS analysis. Five μL of sample was injected into the LTQ Orbitrap Velos mass
spectrometer with Dionex U-3000 Rapid Separation nano LC system (Thermo Scientific) for analysis. MS data were acquired automatically using Analyst QS 1.1 software (Applied Biosystems)
following a MS survey scan over m/z 350–1500 at a resolution of 60,000 for full scan and 2, 000 for MS/MS measurements. The LC-MS/MS data were searched against the _P. fucata_ predicted
protein database (http://marinegenomics.oist.jp/genomes) using a Mascot 2.1 search engine with carbamidoethylated cysteine as a fixed modification and oxidized methionine and tryptophan as
variable modifications. The peptide MS and MS/MS tolerances were set to 0.5 Da. Finally, sequences with mascot scores of at least 5.0 and with at least two matched peptide fragments were
considered valid. NUCLEOTIDE AND AMINO ACID SEQUENCES ANALYSIS Identification of proteins from above was attempted using Blastp and tBlastn searches against NCBI database
(http://blast.ncbi.nlm.nih.gov/Blast.cgi). The protein sequences were deposited in NCBI (Table S1).The theoretical mass, isoelectric point and amino acid composition of the proteins were
computed using ProtParam from the EXPASY online server. Conserved domains were predicted using SMART (http://smart.embl-heidelberg.de/) and InterproScan (http://www.ebi.ac.uk/interpro/
search/sequence-search). IUPRED (http://iupred.enzim.hu/) was used to recognize disordered regions from the amino acid sequences of SMPs based on the estimated pairwise energy content.
XSTREAM (http://jimcooperlab.mcdb.ucsb.edu/xstream/) was used to isolate proteins with tandem-arranged repeat units using the default settings. QUANTIFICATION OF NACRE AND PRISM TRANSCRIPTS
BY REAL-TIME PCR Total RNA from the mantle tissue and muscle were extracted using TRIzol reagent (Life Technologies) according to the manufacturer's instructions. RNA integrity was
checked by agarose gel analysis and RNA concentrations were determined by NanoDrop 2000 (Thermo Scientific). RNA concentration was approximately 1000 ng·μL−1 and A260/A280 was above 1.90.
Then, cDNA was prepared by reverse transcription-PCR of the total RNA with GoScriptTM Reverse Transcription System (Promega) following the manufacturer’s instructions. Real-time PCR was
conducted to quantify gene expression levels, with β-actin as an internal reference due to its relatively stable expression in mantle cells15,16. A typical reaction mixture is: SYBR® Premix
Ex TaqTM II(Takara) 12.5 μL, forward primer 0.5 μL (10 μM), reverse primer 0.5 μL (10 μM), cDNA template 0.5 μL and H2O 6 μL. PCR parameters were: 95 °C for 30 s (1 cycle); 95 °C for 5 s, 60
°C for 30 s (40 cycles), 72 °C for 5 s; 72 °C for 30 s (see Table S2 for primers). Dissociation curves were generated to determine product purity and amplification specificity. Relative
gene expression levels were calculated using two reference genes by the delta-delta method, as follows: fold = 2−[ΔCt sample – ΔCt calibrator] = 2−ΔΔCt. Here, the “ΔCt calibrator” represents
the mean ΔCt values of β-actin (AF378128.1) in the corresponding tissues. RESULTS AND DISCUSSION PROTEIN COMPOSITION IN THE PRISMATIC AND NACRE MATRIX The shell of _P. fucata_ is composed
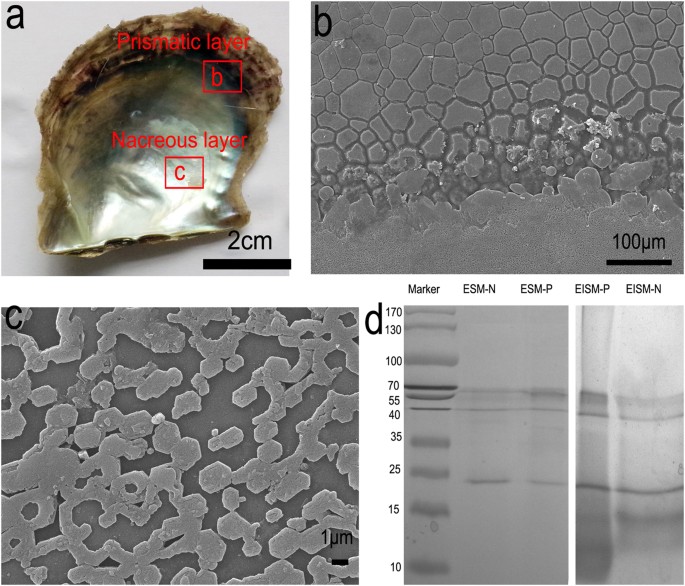
of two layers, the prismatic layer and the nacreous layer (Fig. 1a). The prismatic layer is composed of prisms with length of 10–40 μm embedded in the organic sheath (Fig. 1b) and the
nacreous layer is formed by stacked hexagonal nanotablets with side lengths of 0.5–3 μm (Fig. 1c). To extract SMPs, shells were first cleaned with NaOH to avoid possible contamination from
outside organic matter. Then, separated shells, prism and nacre were dissolved with EDTA, leaving soluble and insoluble extracts. EDTA can chelate Ca2+, dissolve the shell and release the
organic matrices. In this study, we observed that after 60 h, the shell can be fully dissolved. The yields of organic matrices from the shell were approximately 1.5–3.5 mg/g (determined by
the concentration of proteins obtained from certain amounts of shell powders). The soluble and insoluble extracts were subjected to SDS-PAGE (Fig. 1d). Protein bands in the gel were cut and
digested with trypsin. Using LC-MS/MS, peptide fragments were searched against the proteome translated from the draft genome of _P. fucata_27. Consequently, through bioinformatics analysis
such as BLAST, InterproScan and SMART, we identified 72 different SMPs out of 144 whole proteomes, in which 36 and 19 are solely found in the prismatic and the nacreous layers, respectively,
while 17 are found in both layers. It should be noted that these identified proteins are related to the number of lysine and arginine residues available for trypsin cleavage in the protein
sequences29. For example, Aspein and Prisilkin-39 lack trypsin cleavage sites, making them unsuitable for standard proteomic detection29. Jeana L. Drake _et al._ found thirty-six skeletal
organic matrix proteins in the coral, _Stylophora pistillata_. Thirty-one were observed with tryptic digestion, while the remaining five were observed only after proteinase K digestion25.
Therefore, the use of other digestion reagents only slightly increases the number of detected SMPs and proteins found using trypsin as a digestion reagent likely represent most proteins in
the shell. A typical process for LC-MS/MS analysis and protein identification is shown in Figure S1. Intrinsic disorder (ID) refers to segments or to whole proteins that have no fixed 3D
structures, with such disorder sometimes existing in the native state. ID domains are key molecular features that contribute to the formation and function of mollusk nacre; John Evans found
that of 39 mollusk aragonite-associated protein sequences, 100% contain at least one region of intrinsic disorder or unfolding30. This researcher proposed that the intrinsically disordered
domains are important for matrix assembly30. Hence, we used IUPRED to check the 35 unique SMPs found by proteomics. The results showed that 22 out of 35 sequences were predicted to have at
least one region of intrinsic disorder (Table S3). Through XSTREAM, 7 out of 35 sequences were predicted to have tandem repeats (Table S3 and Figure S2). It is noteworthy that repetitive low
complexity domains (RLCDs) are important but not the only implications for intrinsically disordered proteins. According to the blastp results in the National Center for Biotechnology
Information (NCBI) database, SMPs were divided into two groups: proteins with homology (e-value ≤ 10−5) (Table 1) and proteins without homology (e-value > 10−5) (Table 2). Compared to the
shell proteomics of _Pinctada margaritifera_22, a closely related species with _P. fucata_, the numbers of proteins found in both layers were significantly improved. Mpn88, Nacrein,
Nacrein-like, Shematrin-1, Shematrin-2, Shematrin-7, PTyr, PTyr1 and PNU1-9 were found in both layers, suggesting their potential roles in the formation of both layers in _P. fucata_.
Conversely, the proteins in the two layers of _P. margaritifera_ were Nacrein, nacre uncharacterized shell protein (NUSP18) and Shematrin 8, implying that the molecular toolkits responsible
for formation of the prismatic and nacreous layers were extremely different. Sequence alignments from diverse mollusks and metazoans (Figure S3) showed that copper amine oxidase,
peroxiredoxin and chitinase were highly conserved across the metazoa. EGF domain-containing proteins and FN3 domain-containing proteins were highly conserved in the Pinctada family.
IMMUNOLOCALIZATION OF PROTEINS To further validate the SMPs _in vivo_ and _in vitro_, immunolocalization experiments were performed. Western blotting using polyclonal antibodies raised
against the mixed shell matrix proteins of _P. fucata_ showed ETDA-soluble matrices (ESMs) and ETDA-insoluble matrices (EISMs) all reacted with the antibodies (Figure S4).
Immunohistochemical results clearly indicated that the SMPs are located in the mantle pallial and the mantle edge (Fig. 2a1) but showed no signal in the control (Fig. 2a2). Immunogold
observations of the prismatic layer revealed that the antibodies exhibited in both the prismatic tablets (Fig. 2b1) and the chitin layer (Fig. 2b3). In the nacreous layer, antibodies
exhibited a specific signal on nacre and localized in the interlamellar matrix that separated nacre tablets (Fig. 2b4) and in the nacre tablets (Fig. 2b2). In contrast, the control showed no
gold nanoparticle signal (Figure S5). Immunolabeling synthetic calcite was conducted to verify the influence of extracted proteins on the growth of CaCO3. In the control group without the
addition of extracted proteins, no fluorescence signal was observed under the same microscopy settings (Figure S6). By contrast, all four groups with the addition of extracted proteins
exhibited fluorescence signals, indicating they could be occluded in/on the CaCO3. Specifically, SMPs from prismatic layers at approximately 1 μg·mL−1 had no noticeable effect on the
morphology of CaCO3 and were evenly distributed (Fig. 2c1,c2). EDTA-soluble matrix from nacreous layers was concentrated in the center of crystals (Fig. 2c3). EDTA-insoluble matrix from
nacreous layers changed the rhombohedral crystals into 5–10 μm rounded particles. In addition, the fluorescence intensity seemed to be concentrated at the edge of particles (Fig. 2c4). These
results show that SMPs originate from the mantle cells and are finally embedded in the shells. In addition, SMPs can affect the CaCO3 crystallization process. Extensive studies have shown
that SMPs, a single protein or mixed proteins can affect the nucleation, polymorphism and morphology of CaCO319,31. The immune assay using antibodies against the SMPs suggests that SMPs from
different parts of shells execute their distinct roles in CaCO3 crystallization, resulting in being occluded in/on CaCO3 with different patterns. The _in vitro_ crystallization experiments
were conducted under a basic condition (pH 8.0), which is close to the pH of seawater (8.2); therefore, the experiment may provide clues to the mechanism behind _in vivo_ mineralization.
VERIFICATION AND QUANTIFICATION OF MATRIX GENES BY REAL-TIME PCR To verify and quantify the proteins found by our proteomic analysis, real-time PCR was performed. As is known, the mantle
edge is responsible for the formation of the periostracum and the prismatic layer, whereas the mantle pallial enabled the formation of nacreous layer22. Therefore, we examined the relative
gene expression of 21 selected genes, which correspond to the proteins found in the proteomic analysis, in the mantle edge and mantle pallial of _P. fucata_ (Fig. 3a and Table S4). According
to previous studies, six developmental stages have been described across the entire _P. fucata_ life cycle, including descriptions of the fertilized egg, trochophore stage, D-shaped stage,
umbonal stage, juvenile and adult32. The calcium carbonate crystal polymorphisms, the shell layer structure and the expression of SMPs change during these stages32. Almost all SMPs showed a
dramatic increase at the adult stage. For example, the expression level of Pif and Prisilkin-39 in the adult stage is 2116.9 and 119.48 times that of the juvenile stage, respectively32.
Hence, in the present study, all RNA was extracted from the mantle of adult oysters. SMPs encoded by the twenty-one genes are one valine (V)-rich protein (Alveoline-like protein), one
glycine and serine (GS)-rich protein (NU7), one aspartic acid (D)-rich protein (PNU6), one tissue inhibitor of metalloproteinase (PTIMP), one Peroxiredoxin, one Copper amine oxidase, two
Complement control protein (CCP) proteins (PU8 and PU10), one Laminin G protein (NU10), two von Willebrand factor type A (vWA) proteins (PNU4 and PNU5), three chitin-binding proteins (PU8,
PNU1 and NU5), four chitinase (Clp1, Clp3, PNU3 and PU12) and four fibronectin type III (FN3) proteins (PU3, PU5, PU6 and PU15). Among all 21 tested genes, 18 genes (_Alveoline-like_, _PU8_,
_PTIMP_, _PNU3_, _Clp3_, _Copper amine oxidase_, _Clp1, Peroxiredoxin_, _PU15_, _PNU1_, _PU3_, _PU12_, _PU6_, _PU10_, _PNU6_, _NU5_, _PU5_ and _NU10_) were highly expressed in the mantle
edge, mantle pallial, or both referenced to the muscle (Table S4), indicating they were likely to be involved in biomineralization process. _PNU4_, _PNU5_ and _NU7_ genes were neither highly
expressed in mantle edge nor mantle pallial, so they may originate from other cells such as hemocytes33 and ultimately be involved in biomineralization34. Log(ME/MP) is the relative
expression in the mantle edge to the mantle pallial, indicating roles of genes in the formation of the prismatic or nacreous layers. The results showed that most proteins found in the
prismatic layer were highly expressed in the mantle edge (Fig. 3a). Surprisingly, PU10 and PU5 were found in the prismatic layer, but the corresponding genes were more highly expressed in
the mantle pallial (Fig. 3a), suggesting their additional roles in nacreous layer formation. Similarly, previous studies showed that shematrin 5, a prism-related SMP, had much greater
expression in the mantle pallial than in the mantle edge35. Additionally, six well-studied genes were also examined by real-time PCR (Figure 3b). _Nacrein_ and _Pif177_ genes are related to
the formation of the nacreous layer and showed high expression in the mantle pallial (~105 and ~10-fold referenced to the muscle, respectively). _Tyrosinase-1_(_Tyr-1_), _KRMP_,
_Primalin-14_ and _Prisikin-39_ are genes related to the formation of the prismatic layer and exhibited high expression in the mantle edge (~104, ~105, ~104 and ~103-fold referenced to the
muscle, respectively). Therefore, these data confirm that these genes are expressed in the calcifying tissues and the corresponding proteins are embedded in the shells of _P. fucata_.
PUTATIVE FUNCTIONS OF THE SMPS BASED ON DOMAINS To gain insight into the functions of SMPs, analysis of their sequences and domains is required. SMPs involved in biomineralization have
several distinct characteristics25: 1) they are enriched in some specific amino acids such as aspartic acid, glutamic acid, glycine and serine, 2) they have flexible secondary structures and
repeated low complexity domains (RLCDs) and 3) they possess multiple modularity. From well-known SMPs, biomineralization-related proteins contained carbonic anhydrases, chitin-binding,
aragonite-binding, vWA and D-rich domains. According the blast results, some well-studied SMPs have been found in the shells, verifying the effectiveness of our method. In general, the
Shematrin18 and Tyrosinase family11 were among the most abundant proteins in the prismatic layer and others included Nacrein5, Chitinase-like protein 1 (Clp1)22, Clp322, KRMP9,
Alveoline-like protein22, Amylase (GenBank, AGN55420.1), Prismalin-148, Glycine-rich protein 2-like (PGRP2)36, Tissue inhibitor of metalloproteinases (TIMPs)37, Liprin-α protein, PPP-1038,
Mantle protein 10 (PFMG1)39, Mpn8829, cement-like protein (SGMP1)22, Actin (GenBank, ACD99707.1) and Copper amine oxidase22. Nacrein5, Pif17713, N167 and N1940 were enriched in the nacreous
layer and others were methionine-rich nacre protein (MRNP)41, MSI80 (GenBank, BAL45933.1), MSI606, Mpn8829, Actin (GenBank, ACD99707.1), Peroxiredoxin22 and Polyubiquitin42 (Table 1). The
SMPs that have previously been thoroughly characterized will not be discussed in this work. Moreover, we identified some domains that are considered to play significant roles in the
formation of biominerals. These domains include Glyco_hydro_18, Cu2_monooxygen, Chitin-binding domain 2 (ChtBD2), Complement control protein (CCP/SUSHI), Epidermal growth factor-like (EGF),
TIMP and Laminin_G_2/3. Compared with domains found in _P. margaritifera_, Glyco_hydro_20, EF-hand and Kunitz-like domains were not found. Kunitz domains (InterPro accession number,
IPR002223) are the active domains of proteins that inhibit the function of protein degrading enzymes. Notably, the proteins extracted from prismatic layers carried more diverse domains than
those from nacreous layers. Based on the domains, the SMPs were classified into the following three groups: (1) PROTEINS THAT POTENTIALLY REGULATE THE EXTRACELLULAR MICROENVIRONMENT:
CARBONIC ANHYDRASE, CHITINASE, CHITIN-BINDING PROTEINS AND TIMP The microenvironment, the immediate small-scale environment of mantle cells and organs responsible for the formation of
shells, includes pH, framework (mostly chitin) and proteinase in the seawater. Therefore, the extracellular microenvironment is critical for shell formation. Three types of proteins are
related to the microenvironment: carbonic anhydrase, chitin relevant proteins and proteinase. Carbonic anhydrase is responsible for controlling pH by converting CO2 to HCO3- and is found in
Nacrein. Chitin is the major framework in which CaCO3 grows20. Four proteins (PClp1, PClp3, PNU3 and PU2) contain Glyco_hydro_18 domains (IPR001223), which belong to a family of glycoside
hydrolases, hydrolyzing the glycosidic bond between carbohydrates or between a carbohydrate and a non-carbohydrate moiety. Specifically, chitinase hydrolyzes chitin oligosaccharides.
Real-time PCR showed that mRNAs of _PClp1_, _PClp3_, _PNU3_ had high expression levels in the mantle edge and mantle pallial, suggesting their critical roles in shell formation (Fig. 3a and
Table S4). A very recent microarray study had shown that chitinases were highly expressed at the D-shaped stage of _P. fucata_ when the shell were first formed32. In addition to chitinase,
another domain related to chitin is chitin-binding domain_2 (IPR002557). This domain was found in four proteins (PNU1, PU8, Pif177 and NU5), suggesting that chitin-binding ability is
required by both layers. _PNU1_ and _PU8_ had higher mRNA expression levels in the mantle edge and lower levels in the mantle pallial and _N-U5_ was the opposite. _Pif177_, an important gene
in the formation of nacre, had higher mRNA expression levels in the mantle pallial and lower in the mantle edge (Fig. 3b). In complex seawater, all secreted SMPs faced degradation by
microorganism proteinases. Action must be taken to address this problem. We found tissue inhibitors of metalloproteinase (PTIMP and PTIMP3), which may complex with extracellular matrix
metalloproteinases such as collagenases and irreversibly inactivate them (IPR001820). Members of this family are commonly found in the extracellular regions of vertebrate species. It has
been inferred that the function of TIMP in _P. martensii_ on nacre formation is to inhibit matrix metalloproteinases (MMP) activity due to their capacity to degrade most components of the
extracellular matrix37. Thus, the existence of TIMP may ensure a “safe” place for other SMPs to execute their functions by creating suitable MMP-to-TIMP ratios. In humans, inhibition of MMP
activity occurs in a 1:1 stoichiometric relationship and an imbalanced MMP to TIMP ratio may lead to various diseases43. In _P. martensii_, knocking down of TIMP by RNA interference results
in abnormal nacre formation. However, a “suitable” MMP-TIMP ratio in the nacre formation has yet unknown. In fact, water-soluble extracts from the nacre of _P. margaritifera_ possessed
proteinase inhibitory activity against proteinase K44. Moreover, TIMP is also found in the shells of _C. gigas_45 and _M. coruscus_23. However, other protease inhibitors, such as Kunitz-like
protease inhibitor or WAP (Whey acidic protein) domains proteins46, are not found in the shells of _P. fucata_. There is also a possibility that the amount of proteinase inhibitors in the
shell are too low to detect. The problem may be addressed by RNA-seq of mantle cells in the future. (2) EXTRACELLULAR MATRIX- (ECM) RELATED PROTEINS: FIBRONECTIN-RELATED PROTEINS, LAMININ
PROTEINS, EGF PROTEINS AND LIPRIN-Α PROTEINS The shells are constantly growing and can be repaired after injury, indicating the possibility of communication between cells in different
systems of _P.fucata_ and the extracellular matrix (ECM). Proteomic data from _C. gigas_ indicated that oyster shell matrix was not formed simply by self-assembling silk-like proteins but by
diverse proteins through complex assembly and modification processes that may involve hemocyte and exosomes45. Indeed, it has been discovered that except RNA transport, ECM-receptor
proteins were the most abundant in shell proteins. In agreement with previous results, we identified four ECM-related domains (fibronectin-related, laminin, EGF proteins and liprin-α) in the
SMPs that may mediate communication between cells and the extracellular matrix. Five uncharacterized proteins from the prismatic layer, PU3, PU5, PU6, PU15 and PU16, contain fibronectin
type III (FN3) repeat regions (Fig. 3a and Table S4). Strikingly, PU3, PU5 and PU6 are among the most expressed proteins in the mantle pallial (Table S4), although they are from the
prismatic layer. PU3, PU5, PU6, PU15 are highly expressed in both mantle edge and mantle pallial, implying their vital roles in the formation of both layers. FN3 is an approximately 100
amino acid domain that contains different tandem repeats with binding sites for DNA, heparin and the cell surface. The majorities of proteins containing FN3 are involved in cell surface
binding or are receptor protein tyrosine kinases or cytokine receptors (IPR003961). The beta-sandwich structure of FN3 closely resembles that of immunoglobulin domains47. Notably, in _C.
gigas_, a gene coding for a fibronectin-like protein was highly expressed at the early developmental stage when larval shells are formed in unison with chitin synthase45. Proteins containing
fibronectin have been found in the shells of various biomineralization species, including _P. margaritifera_22, _C. gigas_45, _M. coruscus_23 and _A. millepora_24, suggesting important but
yet unknown roles. PU8 and NU10 possess a Laminin_G (LG) domain, which is thought to mediate attachment, migration and organization of cells into tissues during embryonic development by
interacting with other extracellular matrix components (IPR001791). This domain has approximately 180–200 residues and is found in many extracellular and receptor proteins. LG modules have
been implicated in interactions with cellular receptors such us α6β1 integrins, sulfated carbohydrates and other extracellular ligands48. In shells, proteins containing Laminin_G domain are
found in _C. gigas_45, _M. coruscus_23 and _L. gigantea_21. PU12 contains two epidermal growth factor-like domains (EGF), which are commonly found in extracellular proteins (IPR000742). EGF
exhibits six conserved cysteine residues linked through three sulfide bonds. These domains are related to the immune system, apoptosis and Ca2+ binding. In shells, EGF proteins are found in
_P. margaritifera_23_, C. gigas_45, _M. coruscus_23 and _L. gigantea_21. Another protein belonging to this category is PLiprin-α, a member of the leukocyte common antigen-related (LAR)
protein tyrosine phosphatase-interacting protein family. This protein binds to the tyrosine phosphatase LAR and appears to localize LAR to cell focal adhesions. This interaction may regulate
the disassembly of focal adhesion and thus help orchestrate cell-matrix interactions (IPR029515)49. However, this protein is only found in the shells of _C. gigas_45 and not in the other
species. Interestingly, FN3, Laminin_G, EGF and Liprin-α proteins are able to interact with integrins, which are transmembrane receptors for cell-cell and cell-ECM interactions50. Integrin
was identified in the matrix proteins of _S. pistillata_, a biomineralization model of coral species25. Moreover, αvβ6 integrin has been shown to be expressed by ameloblasts and plays a
crucial role in regulating amelogenin deposition and enamel biomineralization51. These studies suggest the integrin has a strong relationship with biomineralizaiton. Indeed, through
real-time PCR, gene expression of _Integrin_ in the mantle of _P. fucata_ is higher than in the gonad, foot, muscle and gill (Figure S7). (3) OTHER PROTEINS OF INTEREST Except for the above
two groups of proteins, some genes are important due to their high expression in the mantle. An acidic protein, four VwA proteins, three complement control protein (CCP) proteins, a V-rich
protein and two enzymes (Tyrosinase and Copper amine oxidase) are discussed. PNU6 is an extremely acidic protein with pI of 3.51 and possesses poly(D)52. The unique primary sequence
indicates its role in the CaCO3 formation, possibly anchoring Ca2+ through the polyD domain, increasing concentration of local Ca2+ and favoring calcite precipitation. The function of PNU6
may be similar to the polyD-containing protein Aspein, an unusually acidic matrix protein found in _P. fucata_17. Protein-protein interactions are important for shell formation because
framework proteins, acidic proteins and others must cooperate to fulfill the complex requirement of biomineralization. VwA domains (IPR002035) in extracellular eukaryotic proteins mediate
adhesion _via_ metal ion-dependent adhesion sites (MIDAS), which are found in Pif177, PNU4, PNU5 and PU4. Three out of four proteins are found in the two layers, implying the importance of
protein-protein interaction in both layers. In shells, vWA proteins are found in _P. margaritifera_22, _C. gigas_45, _M. coruscus_23 and _L. gigantea_21. The complement control protein (CCP)
modules (also known as short consensus repeats SCRs or SUSHI repeats) contain approximately 60 amino acid residues. They exist in a wide variety of complement and adhesion protein52 and are
found in PU4, PU8 and PU10. Some of the proteins in this group are responsible for the molecular basis of the blood group antigens, which are surface markers on the outside of the red blood
cell membrane (IPR000436). The CCP proteins indicate a putative relationship between shells and the immune system. CCP proteins are found in the shells of _P. margaritifera_22, _C.
gigas_45, _M. coruscus_23 and _S. pistillata_25. Alveoline-like protein (Alv), a V-rich protein, has only been discovered in the other two related species _P. margaritifera_ and _P.
maxima_22. Real-time PCR shows that it is highly expressed (almost 103–104 times referenced to the muscle) in both the mantle edge and mantle pallial (Fig. 3a and Table S4), indicating its
critical roles in the formation of both calcite and aragonite. However, the function of Alv in CaCO3 crystallization is poorly understood. Tyrosinase is an oxidase that controls the
production of melanin by hydroxylation of a monophenol to o-quinone (IPR002227). It is reported that 21 tyrosinase genes were found in the genome of _P. fucata_53. As expected, four
tyrosinase proteins were identified in the prismatic layer and two tyrosinase proteins were found in both layers. Tyrosinase is also found in the shells of _P. margaritifera_22, _C. gigas_45
and _M. coruscus_23. Although the specific roles of tyrosinase are unknown, it is deduced that this protein plays distinctive roles in melanogenesis in pigmented shells11. Additionally, a
tyrosinase gene is potentially involved in larval shell biogenesis in _C. gigas_54. Copper amine oxidase catalyzes the oxidation of a wide range of biogenic amines including
neurotransmitters, histamine and xenobiotic amines (IPR000269). In eukaryotes, they have a broad range of functions including cell differentiation and growth, wound healing, detoxification
and cell signaling. In _P. fucata_, copper amine oxidase is primarily expressed in the mantle edge and has lower levels in the mantle pallial. However, its role has not previously been
investigated in mollusks. A previous study in eastern oysters showed that the amine metabolic process was enriched in SMPs by Gene Ontology enrichment analysis34. This protein has not been
reported in other shells of mollusks. THE INDICATION OF PROTEOMICS ON THE SHELL MINERALIZATION MECHANISM From our proteomic findings, some unexpected proteins have been discovered,
indicating the intricacy of biomineralizaion in pearl oyster. The increased SMPs offer a chance to refine the previously proposed “chitin-silk fibroin gel proteins-acidic macromolecules”
model20. The biomineralization mechanism of nacre has been previously proposed to consist of the following four stages: (1) assembly of the matrix, (2) the first-formed mineral phase, (3)
nucleation of individual aragonite tablets and (4) growth of the tablets to form the mature tissue20. At the first stage, the matrix is formed by layers of β-chitin, with a gel comprising
silk-like protein filling the space between. Using proteomics, we identified four chitinases in both layers of shells. Chitinase is an enzyme that catalyzes the hydrolysis of
β-1,4-N-acetyl-d-glucosamine linkages in chitin polymers and oligomers. Interestingly, we did not find any chitin synthase in our proteomic analysis, although chitin synthase genes can be
found in the genome of _P. fucata_53. These results indicate that chitin synthases are located in the cell or on the cell membrane, while chitinases are secreted to the outside of cell to
reconstruct the chitin network. Chitin binding proteins are able to interact with both chitin and minerals. For example, Pif protein with chitin domains have been proven to play an important
role in the association of the inorganic phase and polysaccharide template and in the controlled nucleation of the initial mineral phase31. _In vivo_, Pif proteins are suggested to be able
to work with other proteins such as N16, contributing to the formation of the lamellar sheet of nacre13. Silk-like proteins are rich in Gly and Ala or just in Gly. Such proteins include
MSI60, MSI80, Shematrins, PGRP2, PAmylase, KRMP4, SGMP1, NU5 and NU7. Silk-like proteins are found in both the prismatic and nacreous layers, indicating their importance. One function of
silk-like proteins is to act as a mild inhibitor of mineralization20. An _in vitro_ crystallization assay demonstrates that recombinant KRMP3 inhibits the precipitation of CaCO3, affects the
crystal morphology of calcite and inhibits the growth of aragonite _in vitro_ and these results are almost entirely attributed to the lysine-rich region. The Gly/Tyr-rich region of KRMP3
has the capacity to bind chitin55. Then, at the second stage, the first-form mineral phase, which is usually composed of amorphous calcium carbonate (ACC), is formed. ACC has been considered
the precursor of biominerals, which exist in a wide range of living organisms, including nacre56. Mollusks requires high concentrations of Ca2+ and CO32- from the seawater to form ACC.
CO32- is concentrated by carbonic anhydrase such as Nacrein and Ca2+ is concentrated by acidic proteins such as Pif13 and Aspein17. ACC is unstable compared to calcite and therefore needs to
be stabilized by specialized macromolecules such as Pif31. Several acidic proteins are found in the shell, including PNU2 and PNU6. At the third stage, nucleation of individual tablets
begins, requiring some nucleators. Asp-rich proteins are thought to play a role in this stage. The final stage is growth of the tablets to form the mature tissue and the shape of the final
biominerals is thought to be thermodynamically driven57. At this stage, SMPs are incorporated in to the final biominerals. It is noteworthy that some SMPs, such as Pif, play multiple roles
and can function during several stages. Furthermore, copper amine oxidase, peroxiredoxin and tyrosinase could be related to modification of SMPs, contributing to the overall structure and
mechanical properties of shells. Although this model can largely explain biomineralization from a crystal growth perspective, many proteins including FN3 proteins, CCP proteins, EGF proteins
found in our proteomic analysis are not relevant to crystal growth so far. Recently, studies show that eastern oyster, _Crassostrea virginica,_ forms its shell through a series of
coordinated events involving hemocyte cells and ECM58. In fact, primary mantle cell cultures of _P. fucata_ are able to precipitate amorphous calcium carbonate _in vitro_, suggesting the
ability of mantle cells to perform biomineralization and shell formation processes59. Therefore, it is reasonable to hypothesize that the shell formation process is related to ECM-related
proteins secreted by the mantle cells. These ECM-related proteins are part of SMPs and may play multiple roles. In fact, Osteopontin, a highly expressed ECM-related protein in bone that is
glycosylated and enriched in acidic residues, is involved in a number of cellular processes including immune response and apoptosis besides its main role in biomineralization60. Although we
are aware that domains do not necessarily represent the exact function of proteins, the results in the present study are able to guide the further study of the diverse shell matrix proteins,
improving our understanding of biomineralization. CONCLUSION Using a proteomic approach, we identified 72 unique shell matrix proteins (SMPs) in which thirty-six are associated with the
prismatic layer and nineteen are associated with the nacreous layer, while seventeen are associated with both layers. Based on immunohistological localization, these proteins were confirmed
in the mantles, shells and synthetic calcites. In addition to controlling the CaCO3 crystallization process, the shell matrix proteins potentially regulate the extracellular microenvironment
and communication between cells and the extracellular matrix (ECM). Our results increase the knowledge of shell matrix proteins in pearl oysters and offer an opportunity to refine the
conventional “chitin-silk fibroin gel proteins-acidic macromolecules” model. ADDITIONAL INFORMATION HOW TO CITE THIS ARTICLE: Liu, C. _et al._ In-depth proteomic analysis of shell matrix
proteins of _Pinctada_ _fucata_. _Sci. Rep._ 5, 17269; doi: 10.1038/srep17269 (2015). REFERENCES * Simkiss, K. & Wilbur, K. M. Biomineralization, (Elsevier, 2012). * Ling Li & Ortiz,
C. Pervasive nanoscale deformation twinning as a catalyst for eficient energy dissipation in a bioceramic armour. Nat. Mater. 13, 1–7 (2014). Article Google Scholar * Kim, H. et al. The
role of nacreous factors in preventing osteoporotic bone loss through both osteoblast activation and osteoclast inactivation. Biomaterials 33, 7489–7496 (2012). Article CAS Google Scholar
* Falini, G., Albeck, S., Weiner, S. & Addadi, L. Control of aragonite or calcite polymorphism by mollusk shell macromolecules. Science 271, 67–69 (1996). Article ADS Google Scholar
* Miyamoto, H. et al. A carbonic anhydrase from the nacreous layer in oyster pearls. Proc. Natl. Acad. Sci. USA 93, 9657–9660 (1996). Article CAS ADS Google Scholar * Sudo, S. et al.
Structures of mollusc shell framework proteins. Nature 387, 563–564 (1997). Article CAS ADS Google Scholar * Samata, T. et al. A new matrix protein family related to the nacreous layer
formation of _Pinctada fucata_. FEBS Lett. 462, 225–229 (1999). Article CAS Google Scholar * Suzuki, M. et al. Characterization of Prismalin-14, a novel matrix protein from the prismatic
layer of the Japanese pearl oyster (_Pinctada fucata_). Biochem. J. 382, 205–213 (2004). Article CAS Google Scholar * Zhang, C., Xie, L. P., Huang, J., Liu, X. L. & Zhang, R. Q. A
novel matrix protein family participating in the prismatic layer framework formation of pearl oyster, _Pinctada fucata_. Biochem. Bioph. Res.Co. 344, 735–740 (2006). Article CAS Google
Scholar * Tsukamoto, D., Sarashina, I. & Endo, K. Structure and expression of an unusually acidic matrix protein of pearl oyster shells. Biochem. Bioph. Res.Co. 320, 1175–1180 (2004).
Article CAS Google Scholar * Zhang, C., Xie, L., Huang, J., Chen, L. & Zhang, R. A novel putative tyrosinase involved in periostracum formation from the pearl oyster _Pinctada
fucata_. Biochem. Bioph. Res.Co. 342, 632–639 (2006). Article CAS Google Scholar * Yan, Z. et al. N40, a novel nonacidic matrix protein from pearl oyster nacre, facilitates nucleation of
aragonite _in vitro_. Biomacromolecules 8, 3597–3601 (2007). Article CAS Google Scholar * Suzuki, M. et al. An acidic matrix protein, Pif, is a key macromolecule for nacre formation.
Science 325, 1388–1390 (2009). Article CAS ADS Google Scholar * Kong, Y. et al. Cloning and characterization of Prisilkin-39, a novel matrix protein serving a dual role in the prismatic
layer formation from the oyster _Pinctada fucata_. J. Biol. Chem. 284, 10841–10854 (2009). Article CAS Google Scholar * Fang, D. et al. Novel basic protein, PfN23, functions as key
macromolecule during nacre formation. J. Biol. Chem. 287, 15776–15785 (2012). Article CAS Google Scholar * Pan, C. et al. A novel acidic matrix protein, PfN44, stabilizes magnesium
calcite to inhibit the crystallization of aragonite. J. Biol. Chem. 289, 2776–2787 (2014). Article CAS Google Scholar * Takeuchi, T., Sarashina, I., Iijima, M. & Endo, K. _In vitro_
regulation of CaCO3 crystal polymorphism by the highly acidic molluscan shell protein Aspein. FEBS Lett. 582, 591–596 (2008). Article CAS Google Scholar * Yano, M., Nagai, K., Morimoto,
K. & Miyamoto, H. Shematrin: A family of glycine-rich structural proteins in the shell of the pearl oyster _Pinctada fucata_. Comp. Biochem. Physiol. B 144, 254–262 (2006). Article
Google Scholar * Seto, J. et al. Nacre protein sequence compartmentalizes mineral polymorphs in solution. Cryst. Growth Des. 14, 1501–1505 (2014). Article CAS Google Scholar * Addadi,
L., Joester, D., Nudelman, F. & Weiner, S. Mollusk shell formation: a source of new concepts for understanding biomineralization processes. Chem-Eur. J. 12, 980–987 (2006). Article CAS
Google Scholar * Marie, B. et al. The shell-forming proteome of _Lottia gigantea_ reveals both deep conservations and lineage-specific novelties. FEBS J. 280, 214–232 (2013). Article CAS
Google Scholar * Marie, B. et al. Different secretory repertoires control the biomineralization processes of prism and nacre deposition of the pearl oyster shell. Proc. Natl. Acad. Sci.
USA 109, 20986–20991 (2012). Article CAS ADS Google Scholar * Liao, Z. et al. In-depth proteomic analysis of nacre, prism and myostracum of Mytilus shell. J. Proteomics 122, 26–40
(2015). Article CAS Google Scholar * Ramos-Silva, P. et al. The skeletal proteome of the coral _Acropora millepora_: the evolution of calcification by co-option and domain shuffling. Mol.
Biol. Evol. 30, 2099–2112 (2013). Article CAS Google Scholar * Drake, J. L. et al. Proteomic analysis of skeletal organic matrix from the stony coral _Stylophora pistillata_. Proc. Natl.
Acad. Sci. USA 110, 3788–3793 (2013). Article CAS ADS Google Scholar * Mann, K. & Jackson, D. J. Characterization of the pigmented shell-forming proteome of the common grove snail
_Cepaea nemoralis_. BMC Genomics 15, 249 (2014). Article Google Scholar * Takeuchi, T. et al. Draft genome of the pearl oyster _Pinctada fucata_: a platform for understanding bivalve
biology. DNA Res., 19, 117–130 (2012). Article CAS Google Scholar * Seto, J., Zhang, Y., Hamilton, P. & Wilt, F. The localization of occluded matrix proteins in calcareous spicules of
sea urchin larvae. J. Struct. Biol. 148, 123–130 (2004). Article CAS Google Scholar * Berland, S. et al. Coupling proteomics and transcriptomics for the identification of novel and
variant forms of mollusk shell proteins: a study with _P. margaritifera_. ChemBioChem 12, 950–961 (2011). Article CAS Google Scholar * Evans, J. S. Aragonite-associated biomineralization
proteins are disordered and contain interactive motifs. Bioinformatics 28, 3182–3185 (2012). Article CAS Google Scholar * Bahn, S. Y., Jo, B. H., Hwang, B. H., Choi, Y. S. & Cha, H.
J. Role of Pif97 in nacre biomineralization: _in vitro_ characterization of recombinant Pif97 as a framework protein for the association of organic–inorganic layers in nacre. Cryst. Growth
Des. 15, 3666–3673 (2015). Article CAS Google Scholar * Jun Liu et al. Microarray: A global analysis of biomineralization-related gene expression profiles during larval development in the
pearl oyster, Pinctada fucata. BMC Genomics 16, 325 (2015). Article Google Scholar * Mount, A. S., Wheeler, A., Paradkar, R. P. & Snider, D. Hemocyte-mediated shell mineralization in
the eastern oyster. Science 304, 297–300 (2004). Article CAS ADS Google Scholar * Wang, X. et al. Oyster shell proteins originate from multiple organs and their probable transport
pathway to the shell formation front. PLoS One 8, e66522 (2013). Article CAS ADS Google Scholar * Kinoshita, S. et al. Deep sequencing of ESTs from nacreous and prismatic layer producing
tissues and a screen for novel shell formation-related genes in the pearl oyster. PLoS One 6, e21238 (2011). Article CAS ADS Google Scholar * Funabara, D. et al. Novel genes
participating in the formation of prismatic and nacreous layers in the pearl oyster as revealed by their tissue distribution and RNA interference knockdown. PLoS One 9, e84706 (2014).
Article ADS Google Scholar * Yan, F. et al. Tissue inhibitor of metalloproteinase gene from pearl oyster _Pinctada martensii_ participates in nacre formation. Biochem. Bioph. Res.Co. 450,
300–305 (2014). Article CAS Google Scholar * Nakayama, S. et al. Identification and characterization of a matrix protein (PPP-10) in the periostracum of the pearl oyster, _Pinctada
fucata_. FEBS Open Bio 3, 421–427 (2013). Article CAS Google Scholar * Liu, H.-L. et al. Identification and characterization of a biomineralization related gene PFMG1 highly expressed in
the mantle of _Pinctada fucata_. Biochemistry 46, 844–851 (2007). Article CAS Google Scholar * Yano, M., Nagai, K., Morimoto, K. & Miyamoto, H. A novel nacre protein N19 in the pearl
oyster _Pinctada fucata_. Biochem. Bioph. Res.Co. 362, 158–163 (2007). Article CAS Google Scholar * Marie, B. et al. Characterization of MRNP34, a novel methionine-rich nacre protein from
the pearl oysters. Amino Acids 42, 2009–2017 (2012). Article CAS Google Scholar * Fang, D. et al. Ubiquitylation functions in the calcium carbonate biomineralization in the extracellular
matrix. PLoS One 7, e35715 (2012). Article CAS ADS Google Scholar * Tency, I. et al. Imbalances between matrix metalloproteinases (MMPs) and tissue inhibitor of metalloproteinases
(TIMPs) in maternal serum during preterm labor. PLos One 7, e49042 (2012). Article CAS ADS Google Scholar * Bédouet, L. et al. Heterogeneity of proteinase inhibitors in the water-soluble
organic matrix from the oyster nacre. Mar. Biotechnol. 9, 437–449 (2007). Article Google Scholar * Zhang, G. et al. The oyster genome reveals stress adaptation and complexity of shell
formation. Nature 490, 49–54 (2012). Article CAS ADS Google Scholar * Marie, B. et al. Proteomic analysis of the organic matrix of the abalone _Haliotis asinina_ calcified shell.
Proteome Sci. 8, 54 (2010). Article CAS Google Scholar * Hytonen, V. P. & Wehrle-Haller, B. Protein conformation as a regulator of cell-matrix adhesion. Phys.Chem.Chem. Phys. 16,
6342–6357 (2014). Article Google Scholar * Timpl, R. et al. Structure and function of laminin LG modules. Matrix Biol. 19, 309–317 (2000). Article CAS Google Scholar * Serra-Pages, C.
et al. The LAR transmembrane protein tyrosine phosphatase and a coiled-coil LAR-interacting protein co-localize at focal adhesions. EMBO J. 14, 2827 (1995). Article CAS Google Scholar *
Humphries, M. Integrin structure. Biochem. Soc.Trans. 28, 311–339 (2000). Article CAS Google Scholar * Mohazab, L. et al. Critical role for alphavbeta6 integrin in enamel
biomineralization. J. Cell Sci. 126, 732–44 (2013). Article CAS Google Scholar * Norman, D. et al. Three-dimensional structure of a complement control protein module in solution. J. Mol.
Biol. 219, 717–725 (1991). Article CAS Google Scholar * Miyamoto, H. et al. The diversity of shell matrix proteins: Genome-wide investigation of the pearl oyster, Pinctada fucata. Zool.
Sci. 30, 801–816 (2013). Article CAS Google Scholar * Huan, P., Liu, G., Wang, H. & Liu, B. Identification of a tyrosinase gene potentially involved in early larval shell biogenesis
of the Pacific oyster _Crassostrea gigas_. Dev. Genes Evol. 223, 389–394 (2013). Article CAS Google Scholar * Liang, J. et al. Dual roles of the lysine-rich matrix protein (KRMP)-3 in
shell formation of pearl oyster, Pinctada fucata. PLos One 10, e0131868 (2015). Article Google Scholar * Nassif, N. et al. Amorphous layer around aragonite platelets in nacre. Proc. Natl.
Acad. Sci. USA 102, 12653–12655 (2005). Article CAS ADS Google Scholar * Bayerlein, B. et al. Self-similar mesostructure evolution of the growing mollusc shell reminiscent of
thermodynamically driven grain growth. Nat. Mater. 13, 1102–1107 (2014). Article CAS ADS Google Scholar * Johnstone, M. B. et al. Cellular orchestrated biomineralization of crystalline
composites on implant surfaces by the eastern oyster, _Crassostrea virginica_ (Gmelin, 1791). J. Exp. Mar.Biol. Ecol. 463, 8–16 (2015). Article CAS Google Scholar * Xiang, L. et al.
Amorphous calcium carbonate precipitation by cellular biomineralization in mantle cell cultures of _Pinctada fucata_. PLoS One 9, e113150 (2014). Article ADS Google Scholar * Giachelli,
C. M. & Steitz, S. Osteopontin: a versatile regulator of inflammation and biomineralization. Matrix Biol. 19, 615–622 (2000). Article CAS Google Scholar Download references
ACKNOWLEDGEMENTS We gratefully acknowledge the support of National Natural Science Foundation of China Grants 31372502, 31572594, 31172382 and Independent Research Projects of Tsinghua
University Grant 20111080964. Chuang Liu gives special thanks to the center of biomedical analysis, Tsinghua University for their technical assistance. AUTHOR INFORMATION AUTHORS AND
AFFILIATIONS * Institute of Marine Biotechnology, Collaborative Innovation Center of Deep Sea Biology, School of Life Sciences, Tsinghua University, Beijing, 100084, China Chuang Liu, Shiguo
Li, Jingjing Kong, Yangjia Liu, Tianpeng Wang, Liping Xie & Rongqing Zhang * Tsinghua-Peking Joint Center for Life Sciences, School of Life Sciences, Tsinghua University, Beijing,
100084, China Chuang Liu Authors * Chuang Liu View author publications You can also search for this author inPubMed Google Scholar * Shiguo Li View author publications You can also search
for this author inPubMed Google Scholar * Jingjing Kong View author publications You can also search for this author inPubMed Google Scholar * Yangjia Liu View author publications You can
also search for this author inPubMed Google Scholar * Tianpeng Wang View author publications You can also search for this author inPubMed Google Scholar * Liping Xie View author publications
You can also search for this author inPubMed Google Scholar * Rongqing Zhang View author publications You can also search for this author inPubMed Google Scholar CONTRIBUTIONS C.L., L.P.X.
and R.Q.Z. conceived the project. C.L., S.G.L., J.J.K., Y.J.L. and T.P.W. performed the experiment and analyzed the data. C.L., L.P.X. and R.Q.Z. wrote the manuscript. All authors discussed
the results and revised the manuscript. ETHICS DECLARATIONS COMPETING INTERESTS The authors declare no competing financial interests. ELECTRONIC SUPPLEMENTARY MATERIAL SUPPLEMENTARY
INFORMATION RIGHTS AND PERMISSIONS This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included
in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain
permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ Reprints and permissions ABOUT THIS ARTICLE
CITE THIS ARTICLE Liu, C., Li, S., Kong, J. _et al._ In-depth proteomic analysis of shell matrix proteins of _Pinctada fucata_. _Sci Rep_ 5, 17269 (2015). https://doi.org/10.1038/srep17269
Download citation * Received: 15 June 2015 * Accepted: 28 October 2015 * Published: 26 November 2015 * DOI: https://doi.org/10.1038/srep17269 SHARE THIS ARTICLE Anyone you share the
following link with will be able to read this content: Get shareable link Sorry, a shareable link is not currently available for this article. Copy to clipboard Provided by the Springer
Nature SharedIt content-sharing initiative